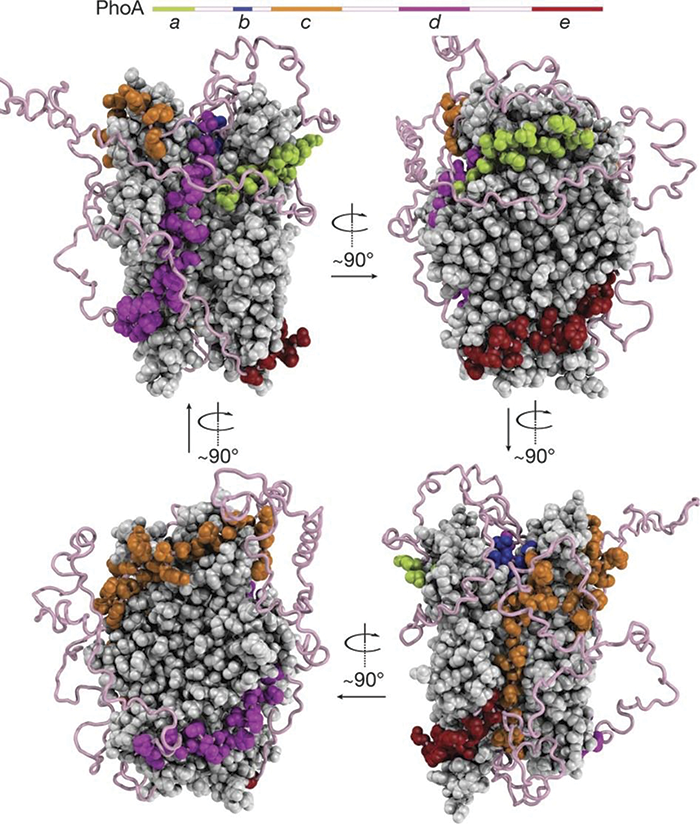
Comparison of SBDβ of BiP–ATP ( deep salmon, PDB 5e84) and BiP–ATP2 ( teal, PDB 6asy) with their superposition based on Cαs of β3–β7. B, overlay of the SBD of BiP–ATP and BiP–ATP2. Ribbon representations of the structures ( insets): ATP-bound BiP with the polypeptide-binding pocket open (BiP-ATP) (PDB 5e84 (8)) ADP-bound BiP from the structures of the isolated NBD (PDB 5evz (102)) SBD (PDB 5e85 (8)) and ATP-bound BiP with the polypeptide-binding pocket fully closed (BiP-ATP2) (PDB 6asy (8)). AMPylated BiP adopts a “domain-docked” structure similar to that of the ATP-bound state even in the apo- or ADP-bound state and is unable to interact productively with ERdjs. Step 6, BiP is post-translationally modified through AMPylation, and this causes the protein to be inactive. Step 5, interaction with ERdjs reorders the polypeptide-binding pocket of the BiP–ATP2 SBD, readying it to interact with another substrate. Step 4, binding of ATP causes a conformational change in the SBD resulting in a more tightly compacted conformation that is thought to “squeeze” the substrate out. 
Step 3, substrate is released with the help of nucleotide-exchange factors (NEFs) that stimulate the release of ADP. This cycle is regulated by ER-localized DnaJ cofactors (ERdjs) that interact with unfolded proteins and transfer them to the ATP-bound form of BiP, while simultaneously triggering ATP hydrolysis. Step 2, upon ATP hydrolysis, the NBD and SBD become undocked, and the lid of the SBD closes providing a form that has high-substrate affinity but slow binding and release rates. Step 1, in the ATP-bound form, the nucleotide-binding domain (NBD) ( blue) and the substrate-binding domain (SBD) ( orange), with its lid open, are docked to each other resulting to a form with high-substrate binding and release kinetics and low-substrate affinity. In this review, we discuss how BiP's functional cycle and interactions with ERdjs enable it to regulate protein homeostasis in the ER and ensure protein quality control.ĪTPase BiP ER-localized DnaJ proteins GRP78 endoplasmic reticulum (ER) endoplasmic reticulum–associated protein degradation (ERAD) heat shock protein (HSP) molecular chaperone protein folding protein misfolding stress response unfolded protein response (UPR).Ī, BiP ATPase cycle. Recent structural and biochemical studies have provided detailed insights into the allosteric regulation of client binding by BiP and have enhanced our understanding of how specific ERdjs enable BiP to perform its many functions in the ER. 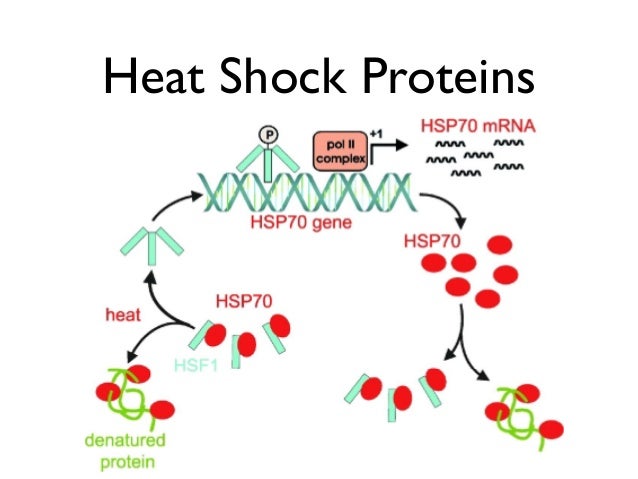 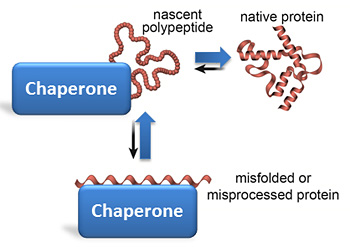
BiP interacts with specific ER-localized DnaJ family members (ERdjs), which stimulate BiP's ATP-dependent substrate interactions, with several ERdjs also binding directly to unfolded protein clients. The ER heat shock protein 70 (Hsp70) family member BiP is an ATP-dependent chaperone that plays a critical role in these processes. An imbalance in protein folding and degradation can result in the accumulation of unfolded proteins in the ER, resulting in the activation of a signaling cascade that restores proper homeostasis in this organelle. Inherent to these processes is a dedicated quality-control system that detects proteins that fail to mature properly and targets them for cytosolic degradation. The endoplasmic reticulum (ER) represents the entry point into the secretory pathway where nascent proteins encounter a specialized environment for their folding and maturation.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
